|
conda warned me about being out of date (although I installed the latest version available from the Anaconda website).I had hoped that Anaconda took automatically care of this matter □ Importing pandas in a notebook gave me complaints about old versions of numpy.Python virtual environments are still some kind of a book of seven seals for me. Packages I like to use in notebooks (such as plotly for data visualization) ended up in other virtual environments than the actual jupyter package, although I never specified a particular environment name.However, I couldn’t get JupyterLab installed with Anaconda this time and wasted several hours scratching my head. In the past, I had good experiences installing Jupyter Notebook using Anaconda. As recommended in the official docs, I went with Anaconda to install the various required Python packages, as I didn’t want to drown in a hot lava sea down in the dependency hell. Until that point it was all fun and games. Tell me, where does it hurt?įinally, I wanted to make the switch and tried to install JupyterLab 1.0.4 based on a Vagrant box ( ubuntu/bionic64). Although the Notebooks app works great, they built a new and more powerful and extensible front end: JupyterLab. The process for Jupyter Notebook is very much the same with one exception the Plot::noteboo_display method must be used to display the plot.Project Jupyter created an indispensable tool for every Data Scientist that finally made exploring and visualizing data a fun thing to do: Jupyter Notebooks. You can also find an example notebook here that will periodically be updated with examples. 
Let trace = Scatter::new(t, y).mode(Mode::Markers) įor Jupyter Lab there are two ways to display a plot in the EvCxR kernel, either have the plot object be in the last line without a semicolon or directly invoke the Plot::lab_display method on it both have the same result. Let t: Vec = linspace(0., 10., n).collect() Then create the following three cells and execute them in order: :dep plotly = UsageĬreate a new notebook and select the Rust kernel. If you're not familiar with the EvCxR kernel it would be good that you at least glance over the EvCxR Jupyter Tour. In a command line execute the following commands: cargo install evcxr_jupyter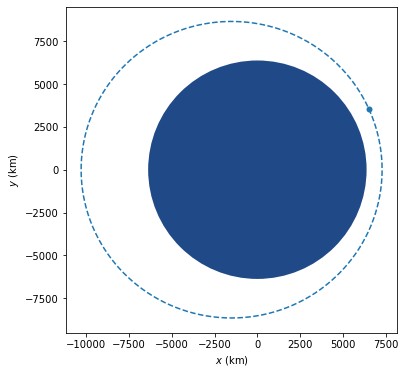 
If CMake is already installed on your system and is in your path (to test that simply run cmake -version if that returns a version you're good to go) then continue to the next steps. Note that EvCxR requires CMake as it has to compile ZMQ. Next you need to install the EvCxR Jupyter Kernel. Open a Python 3 kernel copy/paste the following code in a cell and run it: import aph_objects as goįig = go.Figure(data=go.Bar(x=, y=)) Run the following to install the Plotly Jupyter Lab extension: jupyter labextension install this step is complete to make sure the installation so far was successful, run the following command: jupyter lab This way users know what to expect and also the folks at Plotly have done already most of the heavy lifting to create an extension for Jupyter Lab that works very well. Optionally (or instead of jupyterlab) you can also install Jupyter Notebook: conda install notebookĪlthough there are alternative methods to enable support for the EvCxR Jupyter Kernel, we have elected to keep the requirements consistent with what those of other languages, e.g. conda install -c plotly plotly=4.9.0Ĭonda install jupyterlab "ipywidgets=7.5" If that is not the case you can follow these instructions to get up and running with Anaconda. It is assumed that an installation of the Anaconda Python distribution is already present in the system. Once you've installed the required packages you'll be able to run all the examples shown here as well as all the recipes in Jupyter Lab! Installation 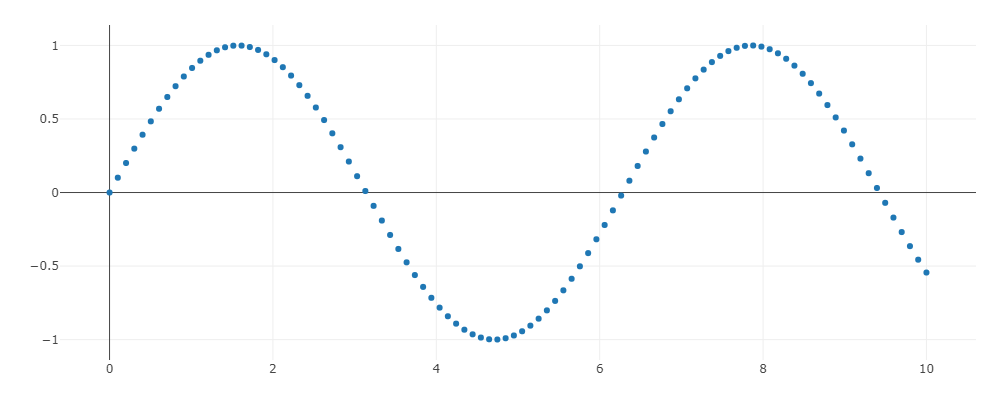
As of version 0.6.0, Plotly.rs has native support for the EvCxR Jupyter Kernel.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
